Machine Learning Software Enhances Molecular Dynamics Modeling
CRD researchers co-author 2020 Gordon Bell Prize Winning Paper
November 19, 2020
Contact: cscomms@lbl.gov
Researchers from Lawrence Berkeley National Laboratory’s Computational Research Division (CRD) are co-authors on a research paper that has been awarded the 2020 ACM Gordon Bell Prize at SC20 for a new machine-learning-based software package that enhances molecular dynamics modeling.
The SC20 paper, “Pushing the limit of molecular dynamics with ab initio accuracy to 100 million atoms with machine learning,” describes DeePMD-kit (DeePMD stands for deep potential molecular dynamics), an optimized ab initio molecular dynamics (AIMD) code developed by an international team that includes Lin Lin, a CRD faculty scientist, and UC Berkeley Mathematics Professor, and Weile Jia, a UC Berkeley postdoc who is a member of Lin’s research group. The project was partially funded by a Department of Energy’s Early Career Research Program award, which Lin received in 2017. Jia and Linfeng Zhang – a research scientist who recently graduated from Princeton University – together led this project.
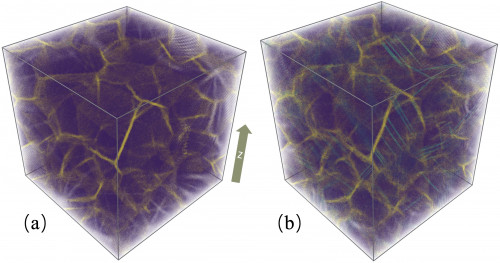
(a) A 10,401,218-atom nanocrystalline copper consisting of 64 randomly oriented crystals with 15-nm averaged grain diameter. (b) The nanocrystalline copper after 10% tensile deformation along the z-axis. Purple, yellow, and cyan denote the atoms in the grains, atoms in the grain boundaries, and atoms in the stacking faults. Credit: Weile Jia, et al
AIMD is a popular simulation tool for studying and describing atomic processes that occur in materials and molecules. While AIMD has long been the method of choice for modeling complex atomistic phenomena from first principles, it relies on computationally expensive electronic structure models such as density functional theory (DFT) and post-DFT theories. Latin for “from the start,” ab initio models calculate interactions between each atom in a model using the most basic laws of physics, sometimes referred to as “first principles calculations.” These models, however, consume an enormous amount of compute power.
“Most DFT-based AIMD calculations are performed with hundreds of atoms for up to a few picoseconds per day,” Lin said. In the past two decades, he added, significant progress has been made in pushing forward the spatial scale of electronic structure methods to study systems comprising up to millions of atoms at least for a certain class of materials. However, relatively little progress has been made in increasing the temporal scale (time scale) accessible to AIMD.
“In application areas such as materials science, chemistry, combustion, and biology-related systems like protein folding, AIMD is the basic tool for understanding atomic structures and atomic behaviors of these systems,” Zhang said. “But all these things are limited by their accuracy when we look at large systems, which current AIMD models cannot handle, even with the help of powerful supercomputers.”
Applying Neural Networks to ab initio Data
To address the grand challenge of modeling larger systems, DeePMD-kit uses a neural network to guide molecular dynamic calculations, Jia explained. By approximating the ab initio data with deep neural networks, DeePMD reduces the computational complexity from cubic to linear scaling, significantly boosting the efficiency. It also demonstrates what can be achieved by integrating physics-based modeling and simulation, machine learning, and efficient implementation on a next-generation computational platform.
Using this approach, DeePMD-kit has the potential to improve the productivity and accuracy of AIMD modeling across chemistry, biology, materials science, and other scientific disciplines, according to the researchers. In early testing, it has been used to model various phenomena in physical chemistry and materials science and demonstrated its ability to boost the spatial and temporal scale accessible by AIMD without losing ab initio accuracy.
For the SC20 paper, the research team ran DeePMD on the Summit supercomputer at the Oak Ridge Leadership Computing Facility (OLCF) on a test case of a block of copper atoms and was able to simulate systems with more than 100 million atoms and efficiently perform AIMD simulations for up to a few nanoseconds per day of supercomputer time. This increases the spatial scale by at least 100x compared to the largest system simulated by all previous Gordon Bell award-winning works, and reduces the time-to-solution by at least 1000x, Lin noted. By efficiently scaling the code up to the entire Summit supercomputer, they attained 91 PFLOPS in double precision (45.5% of the peak) and 162/275 PFLOPS in mixed-single/half precision. “This opens the door to simulating unprecedented size and time scales with ab initio accuracy,” the researchers write.
DeePMD-kit is open source and can be adapted to current and future computing platforms beyond CPUs and GPUs; several research groups are already working on projects related to this, according to Jia. The group is now restructuring DeePMD and merging aspects they modified for the current HPC project so that DeePMD-kit can be useful both for large-scale applications involving systems with 100 million atoms, as well as for applications at smaller scales when only a few GPUs are available. “The latter paradigm may be even more useful for many users,” Jia said.
“The good thing is that most of the techniques we used for thousands of GPUs (on Summit) also work for one or two GPUs, which means the GPU-enabled DeePMD-kit can be applied to a broad range of applications,” Zhang added.
In addition to Jia, Lin, and Zhang, co-authors on the SC20 paper include Han Wang (Institute of Applied Physics and Computational Mathematics, Beijing); Mohan Chen and Denghui Lu (Peking University, Beijing); and Roberto Car and Weinan E (Princeton University)
OLCF is DOE Office of Science user facility.
About Berkeley Lab
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 16 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science.









