HP-CONCORD Paves the Way for Scalable Machine Learning in HPC
Optimized Algorithm Reduces Communication Roadblocks on High Performance Computing Systems
October 1, 2018
Contact: Kathy Kincade, kkincade@lbl.gov, +1 510 495 2124
Some of the most challenging problems in data-driven science involve understanding the interactions between thousands or even millions of variables: how a disease may be caused by a subset of the 20 thousands of human genes, or agricultural production improved by a combination of microbial species among millions in the environment. The problem is to discover the most significant relationships between all of these variables (genes that actively work together), while separating the accidental relationships (genes that occasionally appear together) or confounding effects (two genes that only interact through a common third gene).
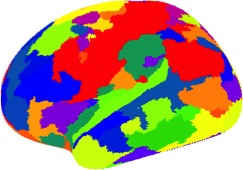
A human brain parcellation derived from HP-CONCORD’s results using a graph clustering algorithm applied to a set of data from the Human Connectome Project.
A powerful machine learning algorithm called CONCORD can identify these relationships, but until recently could only be applied to modest-sized data sets. Researchers from Lawrence Berkeley National Laboratory (Berkeley Lab) and their collaborators have changed that, unleashing the full power of the Department of Energy’s supercomputers on these problems through a high-performance computing version of the algorithm called HP-CONCORD. Using supercomputers at Berkeley Lab’s National Energy Research Scientific Computing Center (NERSC), they demonstrated this parallel algorithm on an enormous set of data from the Human Connectome Project, which computed estimates for about 4 billion parameters, and an even larger demonstration problem with over 800 billion parameters. A paper introducing HP-CONCORD was presented at the 21st International Conference on Artificial Intelligence and Statistics (AISTATS) conference in April 2018.
CONCORD was developed by Sang-Yun Oh, assistant professor in the Department of Statistics and Applied Probability at the University of California, Santa Barbara, as part of his dissertation work at Stanford. Oh was a postdoc at Berkeley Lab when then-graduate student researcher, Penporn Koanantakool, began work on HP-CONCORD as part of her own dissertation at UC Berkeley. CONCORD is an example of a graphical model estimator, a class of machine learning methods that are easier to explain and interpret than some of the competing methods that act more like black boxes. In order to use very large data sets, Koanantakool brought in her perspective on how to make parallel algorithms run across thousands of computational nodes by reducing the amount of communication.
Parallel Scaling via Communication Avoidance
“The most expensive thing you do on any computer is move data around, so you want to minimize data movement between a processor and its own memory and between multiple processors on a parallel machine,” said Kathy Yelick, Associate Lab Director for Computing Sciences at Berkeley Lab and Koanantakool’s thesis advisor. “Reducing data movement tends to save both time and energy.”
Koanantakool, who now works at Google Brain, developed HP-CONCORD and the underlying communication-avoiding algorithms for parallelizing some of the most challenging “all-to-all” style computations.
“When computing the forces between all pairs of particles, or multiplying two matrices, there is a pattern of taking all combinations of things, which involves a lot of communication on a parallel machine,” she explained. Within HP-CONCORD she looks at the problem of multiplying a huge sparse matrix (mostly zeros) with a smaller dense one, which has the added complexity of dividing the nonzeros and computational work evenly across the processors. Her work, which includes extensive experiments on NERSC supercomputers, demonstrates that with HP-CONCORD, the communication is minimal. Her algorithm proved to be more 100 times faster than the standard approach when running on 1,536 cores.
Applications in Data-driven Scientific Discovery
In their 2018 AISTATS paper, the HP-CONCORD team used fMRI (functional magnetic resonance imaging) data to estimate the underlying conditional dependency structure of the brain and then use the resulting estimate to automatically identify functional regions of the brain.
“We constructed a huge brain functional connectivity graph with HP-CONCORD. Then, using this graph, we can draw a map of functional regions in the brain, which is something neuroscientists care about,” Aydin Buluç, a scientist at Berkeley Lab and a co-author on the paper. “The fMRI data we used was not big enough to push HP-CONCORD's limits; however, the datasets will only get bigger.”
Many other science areas will benefit from HP-CONCORD, he emphasized, such as trying to figure out if a trait of a plant is correlated with factors like soil composition, the amount of sunlight it absorbs and its genetic makeup - “all different kinds of objects and variables,” said Buluç.
In statistical terms, HP-CONCORD estimates the most significant parameters in the inverse covariance matrix. Capturing these parameters results in a sparse estimate, which is shown to have good statistical properties when the number of data points is small relative to the number of features, as is most likely the case in many high dimensional datasets.
“Inverse covariance estimates have many practical uses, including reconstructing gene regulatory networks in biology, capturing volatility structure in finance, estimating temperature-to-environmental-proxy relationship in environmental sciences. HP-CONCORD solutions can be used for hypothesis generation in exploratory data analysis to guide further experimental study,” said Oh. “Also, HP-CONCORD estimates can be used as plug-in estimates when relative magnitudes of associations are needed for some downstream analysis.”
Other co-authors of the paper include Alnur Ali at Carnegie Mellon University and Ariful Azad, Dmitriy Morozov and Leonid Oliker from the Computational Research Division at Berkeley Lab.
NERSC is a DOE Office of Science User Facility.
Availability of HP-CONCORD Software
HP-CONCORD and the underlying sparse-dense matrix routines are publicly available on Bitbucket. These are also provided as a ready-to-use software module on NERSC systems. For more details, see the instructions on the Bitbucket page.
About Berkeley Lab
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 16 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science.









