NWB:N Consortium Rolls Out v2.0 of Software Ecosystem for Neuroscientists
February 4, 2019
The Neurodata Without Borders: Neurophysiology (NWB:N) project, led by Berkeley Lab, has achieved yet another milestone with the full release of NWB:N 2.0 in January. Originally launched in 2014, NWB:N is an emerging data standard for neurophysiology research, providing neuroscientists with a software ecosystem that enables them to share, archive, use and build tools for analyzing neurophysiology data.
“NWB:N addresses a critical gap in the neurophysiology community by codifying the wealth of metadata associated with diverse experiments required to turn numbers into knowledge,” said Kristofer Bouchard, PI of the Neural Systems and Engineering lab in the Biological Systems and Engineering Division (BSE) and group lead of the Computational Biosciences Group in Berkeley Lab’s Computational Research Division (CRD).
Oliver Rübel, a research scientist in CRD’s Data Analytics and Visualization Group and lead of the NIH-funded NWB:N project, further said, “With NWB:N 2.0, we have made significant advances toward creating a usable standard, software ecosystem and vibrant community for standardizing neurophysiology data. NWB:N 2.0 is more than just a file format; it defines an ecosystem of standards."
NWB:N is a consortium of neuroscience researchers and computer scientists who are working together to standardize neurophysiology data to ensure the success of brain research worldwide and accelerate the pace of scientific discovery. The consortium was initiated by the Kavli Institute in mid-2014 following the National Institutes of Health’s (NIH) announcement of the BRAIN Initiative. Berkeley Lab’s leadership in the NWB:N project evolved out of a Laboratory Directed Research and Development project, BRAINformat: A Data Standardization Framework for Neuroscience Data.
Over the past year, with funding from the Kavli Foundation, Berkeley Lab scientists Oliver Rübel, Andrew Tritt and Kristofer Bouchard led the development of the beta version of NWB:N 2.0; this work has been in close collaboration with UC San Francisco’s Loren Frank and Eddie Chang, Lydia Ng and others from the Allen Institute for Brain Science, the NWB:N Executive Board members and the broader NWB:N community. In November 2018 the consortium announced that it would receive $2 million from NIH to develop a next-generation data format and software ecosystem.
NWB:N 2.0 Features
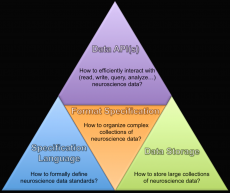
With version 2.0, NWB:N now offers a modern, open software strategy and critical advances in the NWB:N data standard to facilitate usability, performance, extensibility and specificity. The developers also identified, created and separated the fundamental components of the NWB:N ecosystem, including the specification language and core data model, the data storage backend, and data APIs. With PyNWB and MatNWB, this release now includes advanced NWB:N-compliant data APIs for both Python and Matlab. “A modular software infrastructure is necessary for the longevity of the software ecosystem, but creating advanced user APIs for the standard has been fundamental to make the standard accessible to end users,” said Tritt of the Computer Architecture Group and lead software architect of PyNWB.
All resources around the NWB:N data standard are available via neurodatawithoutborders.github.io. Upcoming events to support the rollout of NWB:N 2.0 include a tutorial at Cosyne 2019, March 4, in Cascais, Portugal; the 5th annual BRAIN Initiative investigators meeting, April 11-13 in Washington, D.C. and the NWB:N user days and developer hackathon May 13-16 at the HHMI Janelia Research Campus in Ashburn, VA.
About Berkeley Lab
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 16 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science.