Berkeley Lab Scientists Help Define the Healthy Human Microbiome
Computing, bioinformatics, and microbial ecology resources play key role in mapping our microbial make-up
June 13, 2012
By Dan Krotz
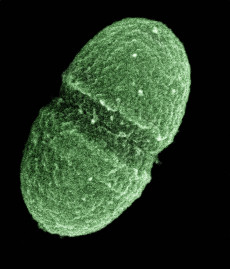
The bacterium, Enterococcus faecalis, which lives in the human gut, is just one type of microbe studied in NIH's Human Microbiome Project. (Courtesy: United States Department of Agriculture)
You’re outnumbered. There are ten times as many microbial cells in you as there are your own cells.
The human microbiome — as scientists call the communities of microorganisms that inhabit your skin, mouth, gut, and other parts of your body by the trillions — plays a fundamental role in keeping you healthy. These communities are also thought to cause disease when they’re perturbed. But our microbiome’s exact function, good and bad, is poorly understood. That could change now that the microbial makeup of healthy humans has been mapped for the first time.
A National Institute of Health (NIH)-organized consortium accomplished this work using software from the Lawrence Berkeley National Laboratory’s (Berkeley Lab's) Computational Research Division, supercomputers at the National Energy Research Scientific Computing Center (NERSC), and networking resources from the Department of Energy's ESnet (Energy Sciences Network).
The research will help scientists understand how our microbiome carries out vital tasks such as supporting our immune system and helping us digest food. It’ll also shed light on our microbiome’s role in diseases such as ulcerative colitis, Crohn’s disease, and psoriasis, to name a few.
In several scientific reports published June 14 in Nature and in journals of the Public Library of Science, about 200 members of the Human Microbiome Project (HMP) Consortium from nearly 80 research institutions report on five years of research.
Berkeley Lab’s role in mapping the human microbiome revolves around big data, both analyzing it and making it available for scientists to use worldwide.
3.5 terabases of data
HMP researchers sampled 242 healthy U.S. volunteers (129 male, 113 female), collecting tissues from 15 body sites in men and 18 body sites in women. Researchers collected up to three samples from each volunteer at sites such as the mouth, nose, skin, and lower intestine. The microbial communities in each body site can be as different as the microbes in the Amazon Rainforest versus the Sahara Desert.
Researchers then purified all human and microbial DNA in more than 5,000 samples and ran them through DNA sequencing machines. The result is about 3.5 terabases of genome sequence data. A terabase is one trillion subunits of DNA.
A comparative analysis system for studying human microbiome samples
Berkeley Lab scientists developed and maintain a comparative analysis system called the Integrated Microbial Genomes and Metagenomes for the Human Microbiome Project (IMG/M HMP). It allows scientists to study the human microbiome samples within the context of reference genomes of individual microbes. Reference genomes help scientists identify the microbes in a sample.
This system is a “data mart” of the larger IMG/M data warehouse that supports the analysis of microbial community genomes at the Department of Energy’s Joint Genome Institute (JGI). IMG/M contains thousands of genomes and metagenome samples with billions of genes. A metagenome consists of the aggregate genomes of all the organisms in a microbial community.
“The IMG/M HMP data mart will help scientists advance our understanding of the human microbiome,” says molecular biologist Nikos Kyrpides of Berkeley Lab’s Genomics Division, who heads the Microbial Genome and Metagenome Programs at JGI. “Scientists can access HMP data with a click of a button and conduct comparative analyses of datasets.”
Kyrpides is also a co-principal investigator of HMP’s Data Analysis and Coordination Center (DACC), together with Victor Markowitz, who heads Berkeley Lab’s Biological Data Management and Technology Center (BDMTC) in the Computational Research Division. Markowitz oversees the development and maintenance of the IMG/M system by BDMTC staff.
“Our system enables scientists worldwide to access and analyze the metagenome datasets generated by NIH’s Human Microbiome Project. We plan to add to our system metagenome datasets generated by similar projects in Europe, Canada and Asia, and thus greatly enhance its comparative analysis potential,” says Markowitz.
A job for high-performance computing
The computation involved in the metagenome data integration underlying IMG/M HMP was partly carried out at the Department of Energy’s National Energy Research Scientific Computing Center (NERSC), which is located at Berkeley Lab. The Energy Sciences Network (ESnet), a high-speed network serving thousands of scientists worldwide that is hosted at Berkeley Lab, was instrumental in transferring the HMP datasets.
Two million computer hours were allocated on NERSC to carry out HMP data integration as well as sift through HMP data for 16S ribosomal RNA genes, which can be used to identify individual species. Focusing on this microbial signature allowed HMP researchers to subtract the human genome sequences and analyze only bacterial DNA.
The analysis helped scientists determine the diversity of microbial species within a person, including within different body sites in a person. It also revealed the extent to which microbial communities vary between people.
“The results suggest that each person has a relatively stable microbiome that is unique to them. You have your own personal microbiome,” says Janet Jansson, a microbial ecologist in Berkeley Lab’s Earth Sciences Division.
In addition, while scientists had previously isolated only a few hundred bacterial species from the body, HMP researchers now calculate that more than 10,000 species occupy the human ecosystem.
What’s next?
“Now that we have a good idea of what makes up the healthy human microbiome, we can study what happens when it’s perturbed because of disease, drugs, or diet,” says Jansson,
In Jansson’s lab, for example, scientists study the role of the gut microbiome in Crohn’s disease, which is an inflammatory bowel disease. Changes in the composition or function of the trillions of microbes inhabiting the human intestine are associated with numerous diseases such as Crohn’s. Understanding the factors underlying these changes will help researchers develop therapies to fight these diseases.
Similar research is also underway at other research centers. Scientists are using HMP data to study the nasal microbiome of children with unexplained fevers. They’re also exploring how the vaginal microbiome undergoes a dramatic shift in bacterial species in preparation for birth, characterized by decreased species diversity.
Other Berkeley Lab researchers with prominent roles in the HMP include Gary Andersen, Shane Canon, and Konstantinos Liolios.
About Berkeley Lab
Founded in 1931 on the belief that the biggest scientific challenges are best addressed by teams, Lawrence Berkeley National Laboratory and its scientists have been recognized with 16 Nobel Prizes. Today, Berkeley Lab researchers develop sustainable energy and environmental solutions, create useful new materials, advance the frontiers of computing, and probe the mysteries of life, matter, and the universe. Scientists from around the world rely on the Lab’s facilities for their own discovery science. Berkeley Lab is a multiprogram national laboratory, managed by the University of California for the U.S. Department of Energy’s Office of Science.
DOE’s Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit energy.gov/science.









